This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Chemical Genetics
Chemical genetics is a research method, used by scientists, to identify small molecules that bind to a specific protein and directly alter its function. Conveniently, when the small molecule is removed, the protein's original function returns. Small molecules refer to small organic compounds chemically produced. To do these experiments scientists isolate a protein of interest, then test a library of small molecules (up to 1 million at a time) for binding to the protein. The small molecules that bind the protein of interest are then retested under different conditions and further tested in cells and animals. These experiments are important to understand what proteins do, how they do it, and to identify small molecules that may be used for medical treatment [1]. Figure 1 shows an example of a small molecule (purple) fitting into the protein 3D-structure and binding. This binding interaction will likely have some effect on the function of the protein.
There are various, untested, small molecules that have the biological process of cell adhesion. Hypothetically speaking, if the mutation in BTBD9 that causes restless leg syndrome does so by disrupting cell adhesion, it is possible that one of these small molecules may be able to aid in treatment by providing the cell adhesion process the protein is missing. The following proteins, for example, might be used to try to treat restless leg syndrome: RGD tripeptide functions in cell adhesion and cell-cell adhesion, K- 252c functions in cell adhesion, and 8- bromo- cAMP functions in cell-matrix adhesion, as does 8- epi- prostaglandin F2alpha. So far, it is not apparent that any chemical genetics screens have been done on BTBD9 and therefore no small molecules have been identified as treatment for restless leg syndrome [2].
There are various, untested, small molecules that have the biological process of cell adhesion. Hypothetically speaking, if the mutation in BTBD9 that causes restless leg syndrome does so by disrupting cell adhesion, it is possible that one of these small molecules may be able to aid in treatment by providing the cell adhesion process the protein is missing. The following proteins, for example, might be used to try to treat restless leg syndrome: RGD tripeptide functions in cell adhesion and cell-cell adhesion, K- 252c functions in cell adhesion, and 8- bromo- cAMP functions in cell-matrix adhesion, as does 8- epi- prostaglandin F2alpha. So far, it is not apparent that any chemical genetics screens have been done on BTBD9 and therefore no small molecules have been identified as treatment for restless leg syndrome [2].
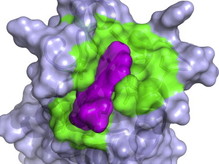
Figure 1: A small molecule, purple, binds into the 3D structure of the protein (green).
RNAi
RNA interference (RNAi) is a research method that is used to silence genes within an organism. This is done by inserting double stranded RNA molecules of a specific gene into the organism. Once inside the organism, the double stranded RNA is diced into fragments. Each fragment then leads a cell to an mRNA that matches its sequence, the cell dices and destroys the mRNA, and the mRNA product is not expressed. Once the gene is silenced, scientists then look for phenotypic changes which they can associate with the silenced gene [3].
Hpo-9, the C.elegans homolog of BTBD9, displays various phenotypes as the result of RNAi experiments. These phenotypes include Bacillus thuringiensis toxin hypersensitive, embryonic lethal, high incidence male progeny, larval arrest, larval lethal, meiosis variant, pore forming toxin hypersensitive, none of which is restless legs (or restlessness in the case of worms) [4]. It is not evident that any RNAi experiments have yet been done on humans or any other model organisms, besides C.elegans.
Hpo-9, the C.elegans homolog of BTBD9, displays various phenotypes as the result of RNAi experiments. These phenotypes include Bacillus thuringiensis toxin hypersensitive, embryonic lethal, high incidence male progeny, larval arrest, larval lethal, meiosis variant, pore forming toxin hypersensitive, none of which is restless legs (or restlessness in the case of worms) [4]. It is not evident that any RNAi experiments have yet been done on humans or any other model organisms, besides C.elegans.
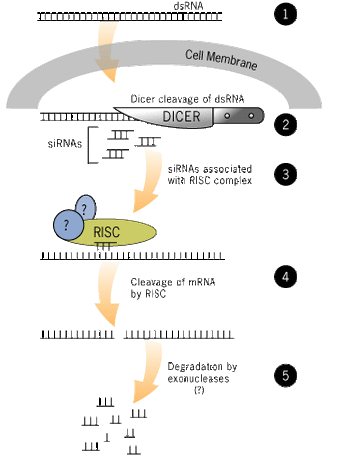
Figure 2: A brief diagram of the steps involved with RNAi. Double stranded DNA is introduced into the cell, it is diced into smaller pieces which bind mRNA, cleave the mRNA, and then degrade the mRNA [8].
Microarray
DNA microarrays are used to measure the expression level of genes within a cell. Thousands of DNA gene sequences are printed onto a glass microscope slide, silicon chip, or nylon membrane. Fluorescently labeled DNA, cDNA, or mRNA is then mixed and hybridized with the DNA spots on the microscope slide. Different colored fluorescent dyes are used when simultaneously comparing the DNA or RNA sequences from multiple samples. From here, a microscope and computer can precisely measure the expression of the genes so that they can be further analyzed [5-6]. Figure 3 shows what a microarray might look like [7].
Although microarray data exists for BTBD9 studies, no microarray studies looking at restless leg syndrome and BTBD9 appear to exist. F
Although microarray data exists for BTBD9 studies, no microarray studies looking at restless leg syndrome and BTBD9 appear to exist. F

Figure 3: What one DNA microarray looks like.
References
[1] http://www.hhmi.org/biointeractive/genomics/poster_a2.html
[2] http://chembank.broadinstitute.org/chemistry/search/execute.htm?id=5660079
[3] http://www.nigms.nih.gov/News/Extras/RNAi/factsheet.htm
[4] http://www.wormbase.org/species/c_elegans/gene/WBGene00015463?query=hpo-9#0a7-9e8-3
[5] Brown, P.O., Botstein, D. (1999). Exploring the new world of the genome with DNA microarrays. Nature America, 21, 33. doi:10.1038/4462
[6] http://www.ncbi.nlm.nih.gov/About/primer/microarrays.html
[7] http://www.glomics.com/s_microservices.html
[8] http://arabidopsis.info/students/rohan/mechanismrnai.html
Image:
http://www.glomics.com/s_microservices.html
[2] http://chembank.broadinstitute.org/chemistry/search/execute.htm?id=5660079
[3] http://www.nigms.nih.gov/News/Extras/RNAi/factsheet.htm
[4] http://www.wormbase.org/species/c_elegans/gene/WBGene00015463?query=hpo-9#0a7-9e8-3
[5] Brown, P.O., Botstein, D. (1999). Exploring the new world of the genome with DNA microarrays. Nature America, 21, 33. doi:10.1038/4462
[6] http://www.ncbi.nlm.nih.gov/About/primer/microarrays.html
[7] http://www.glomics.com/s_microservices.html
[8] http://arabidopsis.info/students/rohan/mechanismrnai.html
Image:
http://www.glomics.com/s_microservices.html
