This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Gene description
BTB (POZ) domain containing 9 (BTBD9) gene has been associated with restless leg syndrome in two genome wide association studies. As of yet, little information is known about the function of BTBD9 in restless leg syndrome, but studies have confirmed a significant correlation with at least five single nucleotide polymorphisms in the DNA [3-4]. A summary of these polymorphisms can be found below. DREME was used to look for DNA motifs, none of which were found [6].
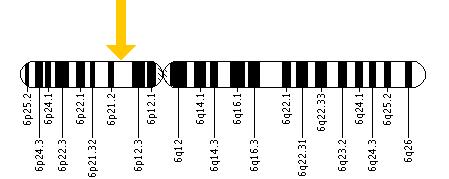
For some specifics, BTBD9 is located at position 21.2 on the short arm of chromosome 6, as is indicated in Figure 1, and is 471,698 bases in length from base pair 38,136,226 to 38,607,923 [1]. The gene has a molecular weight of 2,710,346.71 Daltons [2], the uniprot number is Q96Q07, the accession number is NM_001099272 , (the NM prefix means the accession number is mRNA), and the BTBD9 gene FASTA sequence can be found here [7].
For some specifics, BTBD9 is located at position 21.2 on the short arm of chromosome 6, as is indicated in Figure 1, and is 471,698 bases in length from base pair 38,136,226 to 38,607,923 [1]. The gene has a molecular weight of 2,710,346.71 Daltons [2], the uniprot number is Q96Q07, the accession number is NM_001099272 , (the NM prefix means the accession number is mRNA), and the BTBD9 gene FASTA sequence can be found here [7].
Single nucleotide polymorphisms in BTBD9 change the risk for restless leg syndrome
According to SNPedia.org, the following five single nucleotide polymorphisms (SNP) are located on Chromosome 6, in the BTBD9 gene, and are associated with a risk for restless leg syndrome.
Rs9394492 is at position 38332610, in an intron sequence. People with alleles T:T at this position are the most common with no increased or decreased risk for restless leg syndrome. Alleles C:T at this position results in a 0.76x decreased risk for restless leg syndrome, and C:C is a less than 0.76x decreased risk.
Rs4714156, is at position 38361112, in an intron sequence. Alleles T:T are standard at this position, C:T decreases the risk for restless leg syndrome by 0.61x, and C:C by less than 0.61x.
Rs9296249, is at position 38365841. Alleles T:T are the most common at this position, C:T decreases the risk for restless leg syndrome by 0.62x, and C:C by less than 0.62x. In addition, a study of this SNP in human BTBD9 found it to be significantly associated with low serum ferritin and as we know, restless legs can be a symptom of low iron storage [5].
Rs9357271 is located at the 38365873 base pair position. Alleles T:T at this position is common and does not increase the risk of RLS, C:T at this position decreases the risk of RLS by 0.63x, and C:C at this position by less than 0.63x. This SNP is predicted to be in an intronic sequence. One study of this polymorphism in the human BTBD9 gene found a significant association with restless leg syndrome, meaning that having this SNP provides an increased risk. However, in a familial study, this BTBD9 SNP was not suggestive of restless leg syndrome, so it was determined that this SNP may only be associated with sporadic restless leg sydrome, not familial [4].
Rs3923809 is at position 38440970, in an intron sequence. Alleles G:G at this position is common and doesn't increase the risk of RLS, A:G at this position increases the risk of restless leg syndrome by 1.9x, and A:A at this position increases the risk to greater than 1.9x.
Rs9394492 is at position 38332610, in an intron sequence. People with alleles T:T at this position are the most common with no increased or decreased risk for restless leg syndrome. Alleles C:T at this position results in a 0.76x decreased risk for restless leg syndrome, and C:C is a less than 0.76x decreased risk.
Rs4714156, is at position 38361112, in an intron sequence. Alleles T:T are standard at this position, C:T decreases the risk for restless leg syndrome by 0.61x, and C:C by less than 0.61x.
Rs9296249, is at position 38365841. Alleles T:T are the most common at this position, C:T decreases the risk for restless leg syndrome by 0.62x, and C:C by less than 0.62x. In addition, a study of this SNP in human BTBD9 found it to be significantly associated with low serum ferritin and as we know, restless legs can be a symptom of low iron storage [5].
Rs9357271 is located at the 38365873 base pair position. Alleles T:T at this position is common and does not increase the risk of RLS, C:T at this position decreases the risk of RLS by 0.63x, and C:C at this position by less than 0.63x. This SNP is predicted to be in an intronic sequence. One study of this polymorphism in the human BTBD9 gene found a significant association with restless leg syndrome, meaning that having this SNP provides an increased risk. However, in a familial study, this BTBD9 SNP was not suggestive of restless leg syndrome, so it was determined that this SNP may only be associated with sporadic restless leg sydrome, not familial [4].
Rs3923809 is at position 38440970, in an intron sequence. Alleles G:G at this position is common and doesn't increase the risk of RLS, A:G at this position increases the risk of restless leg syndrome by 1.9x, and A:A at this position increases the risk to greater than 1.9x.
References
[1] http://ghr.nlm.nih.gov/gene/BTBD9
[2] http://www.geneinfinity.org/sms/sms_DNAMW.html
[3] Winkelmann, J., et al. (2007). Genome-wide association study of restless legs syndrome identifies common variants in three genomic regions. Nature Genetics, 39(8), 1000-1006. doi:10.1038/ng2099
[4] Yang, Q., et al. (2011). Association studies of variants in MEIS1, BTBD9, and MAP2K5/SKOR1 with restless legs syndrome in a US population. Sleep Medicine, 12, 800-804. doi:10.1016/j.sleep.2011.06.006
[5] Sorensen, E., (2012). A genetic risk factor for low serum ferritin levels in Danish blood donors. Blood Donors and Blood Collection, 52(12), 2585-2589. doi: 10.1111/j.1537-2995.2012.03629.x
[6] Timothy L. Bailey, "DREME: Motif discovery in transcription factor ChIP-seq data", Bioinformatics, 27(12):1653-1659, 2011.
[7] http://www.ncbi.nlm.nih.gov/projects/RefSeq/key.html
[2] http://www.geneinfinity.org/sms/sms_DNAMW.html
[3] Winkelmann, J., et al. (2007). Genome-wide association study of restless legs syndrome identifies common variants in three genomic regions. Nature Genetics, 39(8), 1000-1006. doi:10.1038/ng2099
[4] Yang, Q., et al. (2011). Association studies of variants in MEIS1, BTBD9, and MAP2K5/SKOR1 with restless legs syndrome in a US population. Sleep Medicine, 12, 800-804. doi:10.1016/j.sleep.2011.06.006
[5] Sorensen, E., (2012). A genetic risk factor for low serum ferritin levels in Danish blood donors. Blood Donors and Blood Collection, 52(12), 2585-2589. doi: 10.1111/j.1537-2995.2012.03629.x
[6] Timothy L. Bailey, "DREME: Motif discovery in transcription factor ChIP-seq data", Bioinformatics, 27(12):1653-1659, 2011.
[7] http://www.ncbi.nlm.nih.gov/projects/RefSeq/key.html