This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
What is Homology?
Homology is a term used to describe a common ancestry when comparing animals [1]. Homology can be examined at various levels of the organism, such as, genes, cells, proteins, tissues, organs, and even behaviors [2]. On this site, I am most concerned about protein homology. A high percentage identity in the RNA sequences means that these proteins are homologous [1]. Shown below are examples of percent identity of the BTBD9 protein for various organisms compared to the human BTBD9 protein. Homologous proteins may further be described as orthologs, meaning the proteins have the same function due to evolutionary speciation, or paralogs, in which the protein homology is a result of sequence duplication that often changes the protein function [1]. It is important to study protein homology to better understand ancestral relationships and how studying model organisms may benefit our knowledge about homologous proteins in humans.
BTBD9 protein homology
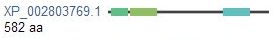
To determine BTBD9 protein homology, Homologene was used to find organisms with a homologous BTBD9 protein; this was determined by aligning the protein sequences of the organisms and doing a BLAST to determine the similarity. BLAST stands for Basic Local Alignment Search Tool and is an algorithm that compares sequences, in this case, protein sequences, to the sequence database [10]. The multiple sequence alignment from Homologene using the program MUSCLE can be seen here. Shown below, in the right hand collumn, are images of conserved protein domains in the homologous BTBD9 proteins, as given by Homologene. Also listed below, is the organism's scientific name and common name, protein name, protein accession number, FASTA nucleotide sequence, E-value, and maximum identity to the human BTBD9 protein, as identified and calculated by Homologene and pairwise alignments in BLAST [3,4,5]. The E-value is the Expectation value, representing the number of different alignments that may occur by chance, so the closer this value is to zero, the more significant the alignment is [10] . Identity is how similar two amino acid sequences are for two aligned proteins [10].
| fastasequences_2.pdf | |
| File Size: | 78 kb |
| File Type: | |
Homologous protein reference numbers

Figure 1: Shown below on the right are conserved domains in homologous BTBD9 proteins. This figure shows the name of the domain and a short description. For the Humans protein domains, in order, are BTB, BACK, and two FA58C domains, this can be used as a reference and guide for reading the homologous proteins thereafter.
Alignments of BTBD9 Homologous Proteins
As explained above, protein sequences were aligned to identify homologous proteins, these sequences were aligned using ClustalW2 and T-Coffee [6-9]. The T-Coffee alignment is in a color which makes it quicker to locate poor alignments. The colors range from red to blue, like a rainbow, with red being a good alignment of the protein sequences and blue being a poor alignment of the protein sequences. The ClustalW2 alignment is also colored but a little more difficult to see areas of poor alignment. The same color at a position indicates alignment. To see the alignments, please download the files below.
|
|
||||||||||||
Analysis and Discussion
Homologous BTBD9 proteins of vertebrates have the highest identity with the human BTBD9 protein, as evident by the high percent maximum identity values above. Furthermore, the low E-values of all the tested organisms above shows the great significance of the protein sequences with human BTBD9. These results describe a high conservation of the BTBD9 protein among organisms, especially vertebrates, implying the importance of this protein's role in the organism. Zebrafish, Fruit Flies, Mosquitos, and Worms (common model organisms) had the lowest identity with the human BTBD9 protein. From these results, Mus musculus, the mouse, would be the best model organism as its protein sequence has a 97% maximum identity to the human protein sequence as well as an E-value of zero.
References
[1] Stéphane Descorps-Declère, Frédéric Lemoine, Quentin Sculo, Olivier Lespinet, Bernard Labedan, The multiple facets of homology and their use in comparative genomics to study the evolution of genes, genomes, and species, Biochimie, Volume 90, Issue 4, April 2008, Pages 595-608, ISSN 0300-9084, 10.1016/j.biochi.2007.09.010. (http://www.sciencedirect.com/science/article/pii/S0300908407002520)
[2] Hall, B. K. (2013), Homology, homoplasy, novelty, and behavior. Dev. Psychobiol., 55: 4–12. doi: 10.1002/dev.21039
[3] http://www.ncbi.nlm.nih.gov/HomoloGene/HGDownload.cgi?hid=14995
[4] Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
[5] Stephen F. Altschul, John C. Wootton, E. Michael Gertz, Richa Agarwala, Aleksandr Morgulis, Alejandro A. Schäffer, and Yi-Kuo Yu (2005) "Protein database searches using compositionally adjusted substitution matrices", FEBS J. 272:5101-5109.
[6] ClustalW and ClustalX version 2 (2007). Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ and Higgins DG. Bioinformatics 2007 23(21): 2947-2948. doi:10.1093/bioinformatics/btm404
[7] A new bioinformatics analysis tools framework at EMBL-EBI (2010). Goujon M, McWilliam H, Li W, Valentin F, Squizzato S, Paern J, Lopez R. Nucleic acids research 2010 Jul, 38 Suppl: W695-9. doi:10.1093/nar/gkq313
[8] T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension.Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W13-7. doi: 10.1093/nar/gkr245. Epub 2011 May 9. PMID: 21558174
[9] T-Coffee: A novel method for fast and accurate multiple sequence alignment. Notredame C, Higgins DG, Heringa J. J Mol Biol. 2000 Sep 8;302(1):205-17. PMID: 10964570
[10] http://www.ncbi.nlm.nih.gov/books/NBK52636/
Header Image: http://www.nature.com/nrg/journal/v8/n6/fig_tab/nrg2099_F2.html
Figures:
http://images.yourdictionary.com/homo-sapiens
http://dogs.about.com/od/fordoglovers/ig/Labrador-Retriever/Duke.-0S-.htm
http://littlefishmag.wordpress.com/2012/11/17/22/funny-cow-28/
http://www.actionbioscience.org/genomic/hhmi.html
http://www.ci.berkeley.ca.us/Health_Human_Services/Environmental_Health/Control___Preventions_of_Rodents.aspx
http://rhamphotheca.tumblr.com/post/14792433381/wild-jungle-fowl-gallus-gallus-by-zoe-gautier
http://www.seymourfish.com/zebra-danio-care/
http://www.taxateca.com/ordendiptera.html
https://koshland-science-museum.org/sites/all/exhibits/exhib_infectious/malaria_vector_control_02.jsp
http://natureafield.com/tag/c-elegans/
http://www.ncbi.nlm.nih.gov/homologene/?term=btbd9&SITE=UniGene%3Acluster.cgi&submit=Go
[2] Hall, B. K. (2013), Homology, homoplasy, novelty, and behavior. Dev. Psychobiol., 55: 4–12. doi: 10.1002/dev.21039
[3] http://www.ncbi.nlm.nih.gov/HomoloGene/HGDownload.cgi?hid=14995
[4] Stephen F. Altschul, Thomas L. Madden, Alejandro A. Schäffer, Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997), "Gapped BLAST and PSI-BLAST: a new generation of protein database search programs", Nucleic Acids Res. 25:3389-3402.
[5] Stephen F. Altschul, John C. Wootton, E. Michael Gertz, Richa Agarwala, Aleksandr Morgulis, Alejandro A. Schäffer, and Yi-Kuo Yu (2005) "Protein database searches using compositionally adjusted substitution matrices", FEBS J. 272:5101-5109.
[6] ClustalW and ClustalX version 2 (2007). Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ and Higgins DG. Bioinformatics 2007 23(21): 2947-2948. doi:10.1093/bioinformatics/btm404
[7] A new bioinformatics analysis tools framework at EMBL-EBI (2010). Goujon M, McWilliam H, Li W, Valentin F, Squizzato S, Paern J, Lopez R. Nucleic acids research 2010 Jul, 38 Suppl: W695-9. doi:10.1093/nar/gkq313
[8] T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension.Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W13-7. doi: 10.1093/nar/gkr245. Epub 2011 May 9. PMID: 21558174
[9] T-Coffee: A novel method for fast and accurate multiple sequence alignment. Notredame C, Higgins DG, Heringa J. J Mol Biol. 2000 Sep 8;302(1):205-17. PMID: 10964570
[10] http://www.ncbi.nlm.nih.gov/books/NBK52636/
Header Image: http://www.nature.com/nrg/journal/v8/n6/fig_tab/nrg2099_F2.html
Figures:
http://images.yourdictionary.com/homo-sapiens
http://dogs.about.com/od/fordoglovers/ig/Labrador-Retriever/Duke.-0S-.htm
http://littlefishmag.wordpress.com/2012/11/17/22/funny-cow-28/
http://www.actionbioscience.org/genomic/hhmi.html
http://www.ci.berkeley.ca.us/Health_Human_Services/Environmental_Health/Control___Preventions_of_Rodents.aspx
http://rhamphotheca.tumblr.com/post/14792433381/wild-jungle-fowl-gallus-gallus-by-zoe-gautier
http://www.seymourfish.com/zebra-danio-care/
http://www.taxateca.com/ordendiptera.html
https://koshland-science-museum.org/sites/all/exhibits/exhib_infectious/malaria_vector_control_02.jsp
http://natureafield.com/tag/c-elegans/
http://www.ncbi.nlm.nih.gov/homologene/?term=btbd9&SITE=UniGene%3Acluster.cgi&submit=Go