This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
BTBD9 protein domains
A protein domain is a segment of the protein that folds separately from the rest of the protein and has it's own function [10]. Pfam and SMART, two databases of protein families and domains, were used to find the BTBD9 protein domains; four domains were found: BTB, BACK and two F5/8 C-type [3]. Figures 1 and 2 show the position and order of the domains within the protein. More information on each domain can be found below.

Figure 1: The location of the BTB/POZ and BACK domains on the BTBD9 human protein as depicted by PFAM [3]. PFAM identified two other domains but did not label them (the blue blocks).
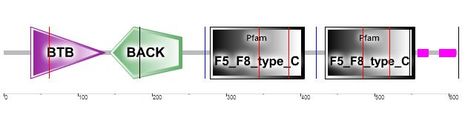
Figure 2: The location of the BTB/POZ and BACK domains in the BTBD9 human protein as depicted by SMART. The two pink regions on the right hand side are areas of low complexity, in other words, areas of ill-defined protein sequence [10].
BTB/POZ domain
The bric-á-brac, tramtrack, broad complex/pox virus and zinc finger domain is more commonly referred to as the BTB/POZ domain. This domain family is under the POZ domain superfamily, as is the BACK domain family [1]. The BTB domain is located near an N-terminal Zinc finger and mediates homomeric dimerization and sometimes heteromeric dimerizations. The function of the BTB/POZ domain in the BTBD9 protein in humans is unknown, although, other proteins with the BTB/POZ domain have been shown to mediate transcriptional regulation, to interact with components of histone deacetylase co-repressor complexes, and to be involved with cytoskeleton dynamics, ion conductance, and protein ubiquitination [2].
BTB domain structure
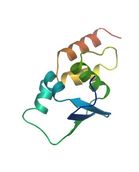
This is an image of the BTB domain folding structure, provided by SWISS-MODEL Repository [4-5, 11]. You can see both alpha helices and beta sheets in this tertiary structure.
BACK domain
BTB and C-terminal Kelch domain, more commonly known as BACK domain, is located next to the BTB/POZ domain containing Kelch repeats (as the name implies). This domain does not have a known function in the BTBD9 protein, but in other proteins it is associated with maintaining substrate orientation, like in E3 ubiquitin-ligase complexes [2].
BTB and BACK domain structures
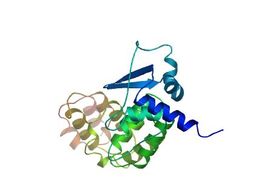
The BTB and BACK domains are located adjacent to each other, pictured here is an image of the BTB and BACK structures together, as depicted by SWISS-MODEL Repository [4-5, 11]. This structure of two adjacent domains is comprised of a mixture of alpha helices and beta sheets.
F5/8 type C Domain
Using SMART, two F5 F8 type C domains were found within the BTBD9 human protein, as shown in Figure 2, although no such domain was described from running the protein sequence through PFAM or numerous other programs. The domain goes by multiple names including Discoidin domain, F5/8 type C domain, or C2-like domain. F5/8 type C domain is described as a C-terminal, twice repeated domain, as you can see two repeats of this domain at the C-terminal end of the sequence in Figure 2. Some extracellular and membrane proteins, as well as blood coagulation factors V and VIII have been shown to contain this F5/8 type C domain. In blood coagulation factors V and VIII this domain participates as part of a larger domain that functions in binding to anionic phospholipids on the surface of platelets and endothelial cells. The second F5/8 type C domain repeat in blood coagulation factor VIII has shown to be involved in the binding and activation of phosphatidylserine. The F5/8 type C domain contains two conserved cysteins in most proteins, which bind through a disulfide bond [7].
F5/8 type C domain structure

This is an image of the F5/8 type C domain structure, from SWISS-MODEL Repository [4-5, 11]. You can see that the structure is a beta barrel composed of eight anti-parallel strands organized in two beta sheets. In the lower section of this beta barrel, there are three loops and numerous basic residues that are very important for lipid binding [7].
Homologous Protein Domains
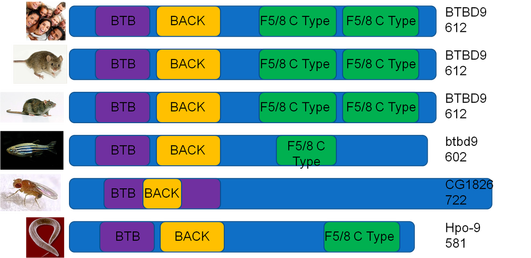
This image compares the domains in homologous BTBD9 proteins. Overall, The BTBD9 protein is well conserved. Humans, mice, and rats have the same amino acid length protein, with the same domains, in the same locations. All shown model organisms have a BTB and BACK domain, potentially suggesting primary importance. On the left is an image of the organism, followed by the protein domains in the center, and the homologous BTBD9 protein name and amino acid length on the right.
Protein isoform domains
There are three protein isoforms as the product of alternative splicing of BTBD9 mRNA. More information on alternative splicing can be found under the Protein homepage.
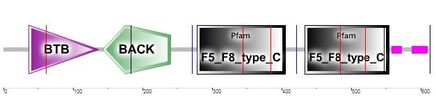
Isoform 1: 612 amino acid length protein. BTB domain position: 36-137. BACK domain position: 143-240. F5/8 type C domain positions: 278-405 and 433-554. The two pink bars are areas of low complexity, positions: 558-571 and 587-608 [6, 8-9].
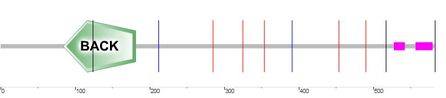
Isoform 2: 582 amino acids in length. Because of alternative splicing, amino acids 1-61 are missing (compared to isoform 1) and the amino acid sequence -MC- is in the first two positions. There is an added 20 amino acid sequqnce at position 383 of isoform 1. There are no BTB or F5/8 type C domains. BACK domain position: 84-181. Low complexity area positions: 528-541 and 557-578 [6, 8-9].
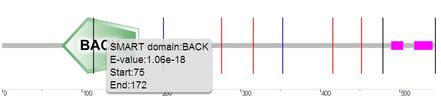
Isoform 3: 544 amino acids in length. Positions 1-68 are spliced out. There are no BTB or F5/8 type C domains. BACK domain position: 75-172. Low complexity areas positions: 490-503 and 519-540 [6, 8-9].
Domain analysis and discussion
The four domains of the BTBD9 protein are very well conserved among the model organisms examined, this suggests the homologous BTBD9 proteins may have similar functions and be allowed for use in research to better understand the human BTBD9 protein. All tested model organisms had a BTB and BACK domain which might suggest the importance of these domains in BTBD9. The three protein isoforms may provide problems in trying to study the BTBD9 protein, particularly the BTB and F5/8 C type domains. These isoforms should be looked at though, in studying restless leg syndrome, as they may be more or less likely to cause restless leg syndrome.
Motifs
A protein motif is a highly conserved region of amino acids within a protein sequence, usually within a protein domain [10]. MEME was used to identify proteins motifs. Three motifs were identified and are shown below along with their E-value, amino acid length, and position within the protein sequence of amino acids [12]. Recall, the lower or close to zero an e-value is, the more significant [13].
Motif analysis and discussionMotif 1 is within the second (more towards the right hand side of the screen) F5/8 C-type domain. Motif 2 is mostly in the second F5/8 C-type domain. Motif 3 is in the first F5/8 C-type domain. No motifs were found in the BTB or BACK domains. This suggests the F5/8 C-type domains are very well conserved.
|
Figure 3: BTBD9 Motif 3.
E-value: 1.1e-405 AA length: 50 Position: 333-383 |
References
[1] http://pfam.sanger.ac.uk/clan/CL0033
[2] http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3334925/
[3] The Pfam protein families database: R.D. Finn, J. Mistry, J. Tate, P. Coggill, A. Heger, J.E. Pollington, O.L. Gavin, P. Gunesekaran, G. Ceric, K. Forslund, L. Holm, E.L. Sonnhammer, S.R. Eddy, A. Bateman
[4] Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009).
The SWISS-MODEL Repository and associated resources.
Nucleic Acids Res. 37, D387-D392.
[5] Jürgen Kopp and Torsten Schwede (2004)
The SWISS-MODEL Repository of annotated three-dimensional protein structure homology models.
Nucleic Acids Res. 32, D230-D234.
[6] http://www.uniprot.org/uniprot/Q96Q07#section_alternative
[7] http://prosite.expasy.org/PDOC00988
[8] Schultz et al. (1998) Proc. Natl. Acad. Sci. USA 95, 5857-5864
[9] Letunic et al. (2012) Nucleic Acids Res , doi:10.1093/nar/gkr931
[10] http://www.ncbi.nlm.nih.gov/books/NBK62051/
[11] Arnold K., Bordoli L., Kopp J., and Schwede T. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22,195-201.
[12] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[13] http://www.ncbi.nlm.nih.gov/books/NBK52636/
Figures:
Domains:
http://swissmodel.expasy.org/repository/?pid=smr01&zid=async
http://smart.embl-heidelberg.de/
http://swissmodel.expasy.org/repository/?pid=smr01&zid=async
Motifs:
http://meme.nbcr.net/meme/
[2] http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3334925/
[3] The Pfam protein families database: R.D. Finn, J. Mistry, J. Tate, P. Coggill, A. Heger, J.E. Pollington, O.L. Gavin, P. Gunesekaran, G. Ceric, K. Forslund, L. Holm, E.L. Sonnhammer, S.R. Eddy, A. Bateman
[4] Kiefer F, Arnold K, Künzli M, Bordoli L, Schwede T (2009).
The SWISS-MODEL Repository and associated resources.
Nucleic Acids Res. 37, D387-D392.
[5] Jürgen Kopp and Torsten Schwede (2004)
The SWISS-MODEL Repository of annotated three-dimensional protein structure homology models.
Nucleic Acids Res. 32, D230-D234.
[6] http://www.uniprot.org/uniprot/Q96Q07#section_alternative
[7] http://prosite.expasy.org/PDOC00988
[8] Schultz et al. (1998) Proc. Natl. Acad. Sci. USA 95, 5857-5864
[9] Letunic et al. (2012) Nucleic Acids Res , doi:10.1093/nar/gkr931
[10] http://www.ncbi.nlm.nih.gov/books/NBK62051/
[11] Arnold K., Bordoli L., Kopp J., and Schwede T. (2006). The SWISS-MODEL Workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22,195-201.
[12] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[13] http://www.ncbi.nlm.nih.gov/books/NBK52636/
Figures:
Domains:
http://swissmodel.expasy.org/repository/?pid=smr01&zid=async
http://smart.embl-heidelberg.de/
http://swissmodel.expasy.org/repository/?pid=smr01&zid=async
Motifs:
http://meme.nbcr.net/meme/



